CUT&RUN Overview
Learn about CUT&RUN—a low cell number alternative to ChIP-qPCR and ChIP-seq to analyze protein-DNA interaction in chromatin.
What is CUT&RUN?
The development of methods that enable mapping of protein-DNA interactions, such as chromatin-immunoprecipitation (ChIP) and ChIP-seq, have led to a growing awareness that aberrant epigenetic regulation drives a wide variety of human diseases. Cleavage Under Targets & Release Using Nuclease (CUT&RUN) is a new technology that can be used for chromatin profiling.
How It Works
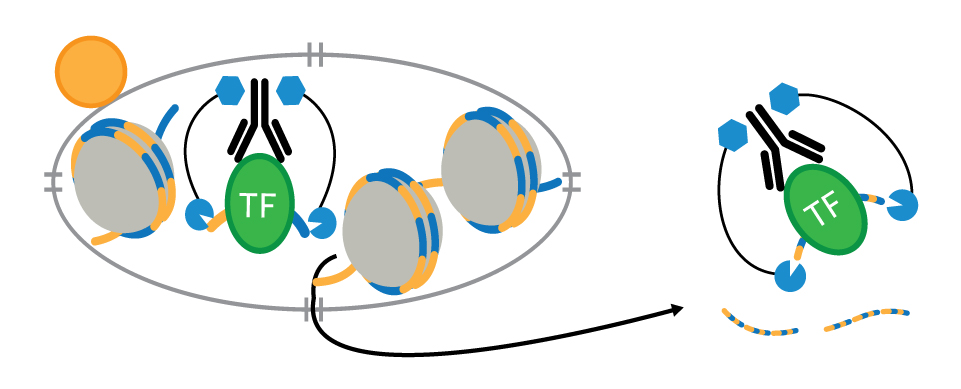
CUT&RUN is an in vivo method that uses a target-specific primary antibody and a Protein A-Protein G-Micrococcal Nuclease (pAG-MNase) to isolate specific protein-DNA complexes1,2,3. It only takes 1 to 2 days to get from cell to DNA and can be automated for maximal throughput and reproducibility4.
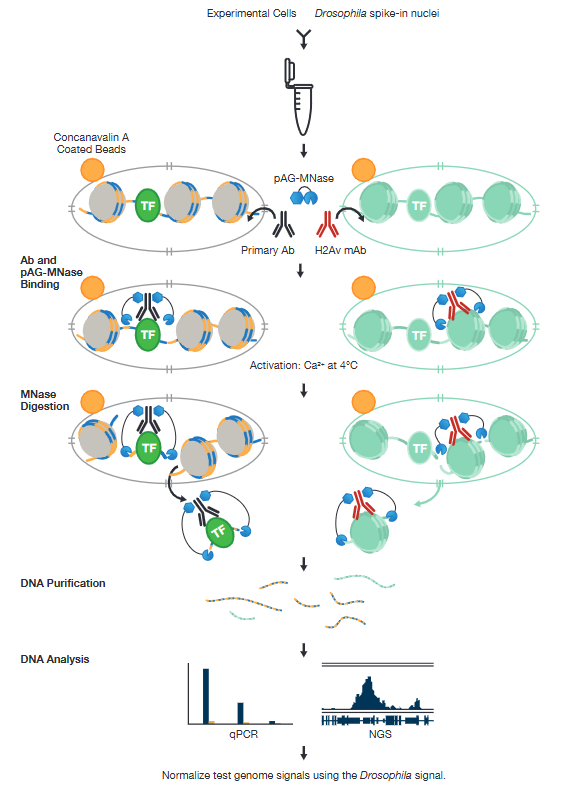
CUT&RUN Workflow Including the Drosophila Spike-in Normalization Strategy
To isolate the protein-DNA complex of interest, cells are first harvested and bound to Concanavalin A-coated magnetic beads to simplify cell handling and minimize cell loss during subsequent washes. Cell membranes are permeabilized with digitonin to facilitate the entry of the primary antibody into the nuclei, where it binds to the histone, transcription factor, or cofactor of interest. The pAG domain of the pAG-MNase fusion protein then binds to the primary antibody heavy chain, targeting the enzyme to the chromatin region of interest. The addition of Ca2+ activates the pAG-MNase to initiate DNA digestion. This allows the cleaved chromatin complex to diffuse away from the genomic chromatin, out of the nuclei, and into the sample supernatant, where it can be collected using either a DNA spin column or phenol/chloroform extraction followed by ethanol precipitation.
The purified, enriched DNA can then be identified and quantified using qPCR or used to construct a DNA sequencing library for next-generation sequencing (NGS) and whole-genome mapping.
CUT&RUN Normalization
To normalize signals across reactions, choose one of two spike-in strategies to make the CUT&RUN assay more quantitative–it would be redundant to employ both.
- For downstream normalization add yeast DNA spike-in after the MNase digestion step.
- For whole-workflow normalization (shown above), add Drosophila spike-in nuclei at the beginning of the experiment; the corresponding H2Av antibody is then added later during the primary antibody incubation step.
Please note that that both spike-in methods are designed for comparing signals across samples targeting the same protein (e.g., comparing H3K27me3 levels across multiple samples). They do not facilitate quantitative comparisons between different targets, such as comparing the signal of an Ezh2 sample to an H3K27me3 sample. Read the corresponding blog to learn more about these spike-strategies.
Choosing the Right CUT&RUN Normalization Control for Your Experiment
CST offers two CUT&RUN kits to meet your specific normalization requirements.
Kit | ||
Control Composition | Drosophila spike-in nuclei and antibody | Yeast spike-in DNA |
Workflow Coverage | Cell handling through permeabilization, antibody binding, chromatin digestion, DNA purification, library prep, and sequencing. | DNA purification, library prep, and sequencing. |
Normalization Focus | Technical variability across the entire assay. | Bias in library prep and sequencing. |
Ideal Use Case | Drug perturbation studies, subtle genome-wide chromatin changes, multi-batch comparisons, and experiments with variable starting material, in addition to variability in library prep and sequencing. | Partial benchmarking and experiments mainly concerned with bias and variability in library prep and sequencing. |
Drosophila Spike-in Normalization Increases Confidence in Drug Response Data
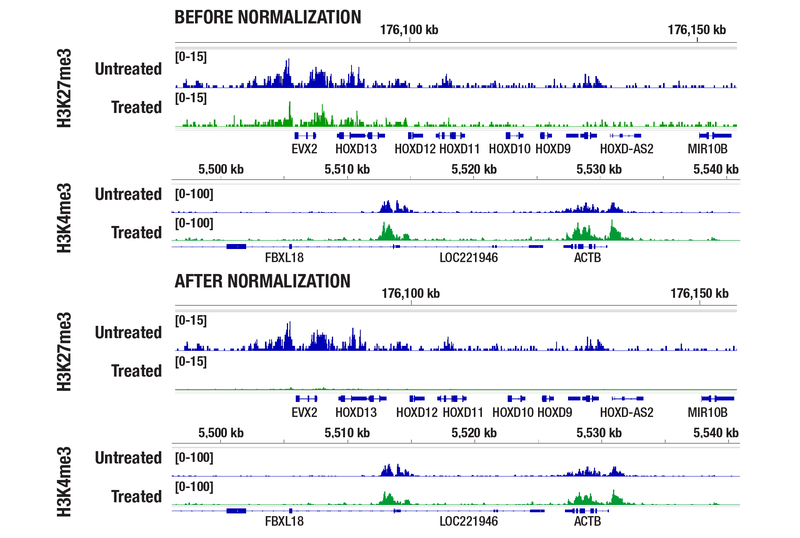
The experiment below demonstrates how the Drosophila spike-in normalization strategy increases confidence that observed changes in global histone marks represent true biological responses, rather than technical variation. In the normalized data, the tazemetostat‑treated sample shows a reduced H3K27me3 signal, consistent with EZH2 inhibition, while H3K4me3 signal remains comparable.
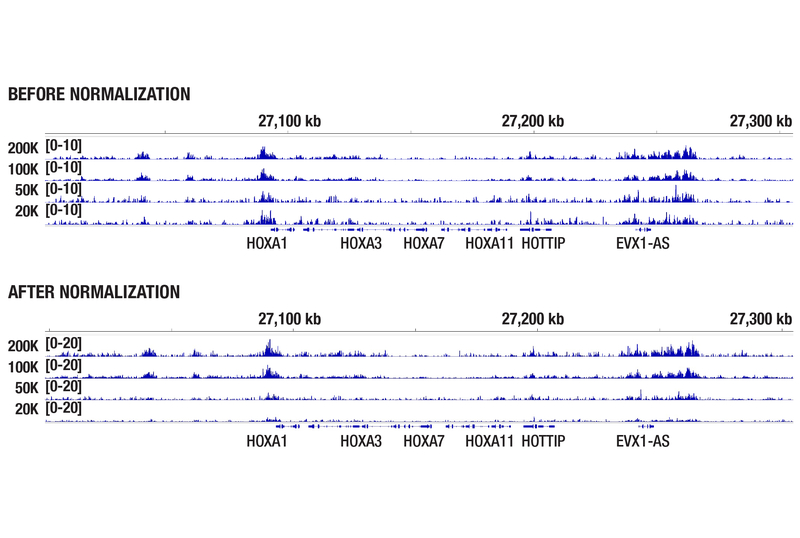
Accurate Signal Normalization Across Cell Titrations with Drosophila Spike-In Control
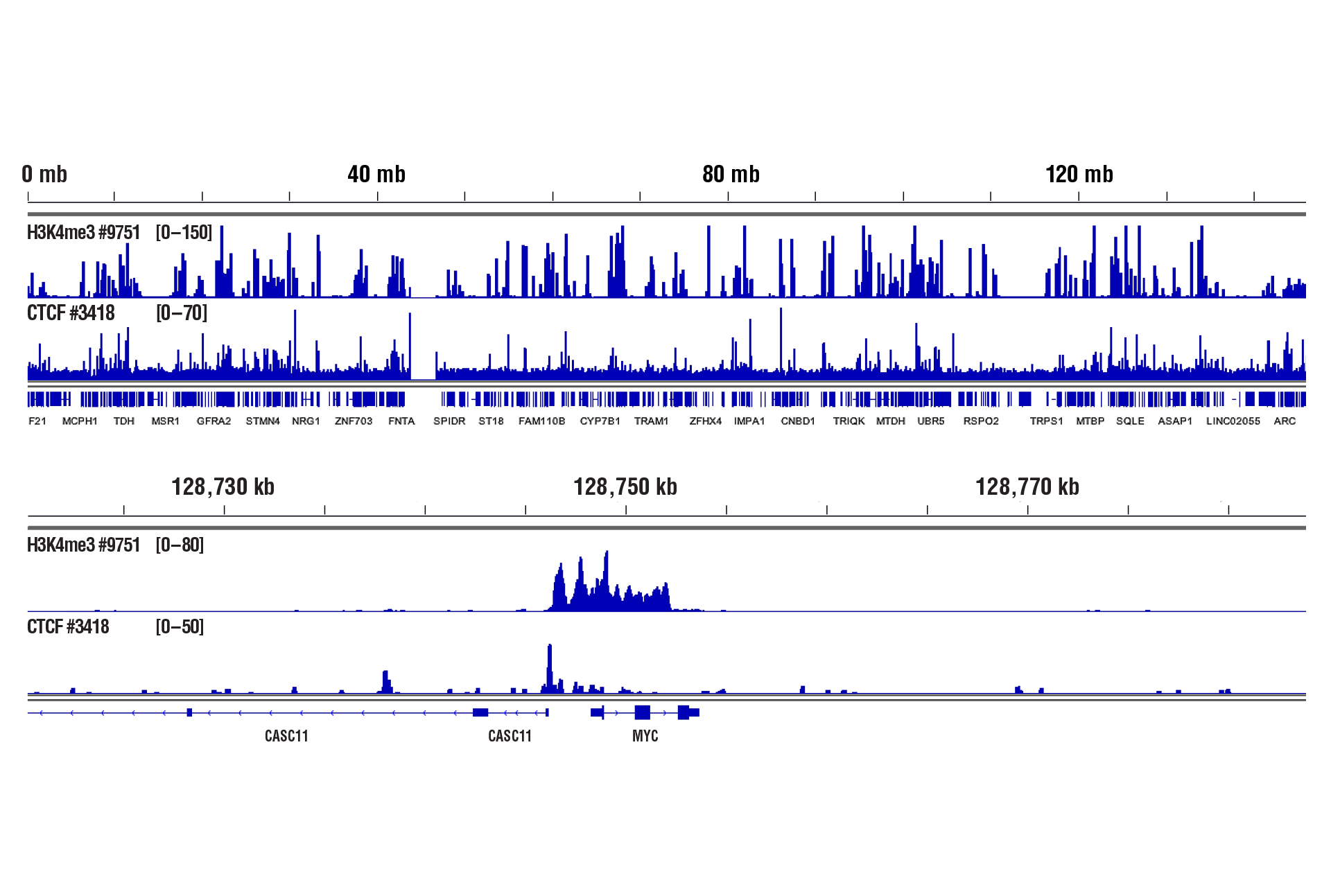
Drosophila spike‑in normalization ensures that CUT&RUN signal intensities scale appropriately with starting cell number. In the cell titration experiment below, this allows for the accurate comparison of H3K27me3 enrichment across the HOXA locus, and increases confidence that observed differences reflect true input‑dependent changes rather than technical variability.
Normalized qPCR Data with Fragmented Yeast DNA
Normalizing your samples with yeast spike-in DNA provides experimental reproducibility between samples within a given experiment or different experiments that are looking at the same protein.
CUT&RUN Benefits
Both CST CUT&RUN kits provide the following benefits:
Fast time to results | 1 to 2 days from cell to DNA |
Refined protocols | Optimized protocols enhance enrichment of low abundance and/or weak binding transcription factors and cofactors. |
Low sample requirement | 100,000 cells is recommended–kits are validated to use with 5,000-10,000 cells for histone modifications and 10,000-20,000 cells for transcription factors and cofactors. |
Compatible with fixed cells | Light fixation keeps cells intact, preserves cell signaling pathways, and enhances the enrichment for accessory components of huge complexes. |
Use fixed or fresh tissue with confidence | Validated and optimized tissue protocols give you data confidence when using ~20X less sample compared to ChIP. |
Study protein-DNA interactions in primary cells | CUT&RUN significantly lowers cell number requirements, making it suitable to use with primary cells. |
Lower sequencing depth = lower sequencing costs | Only requires 3-5 million high quality reads per sample due to the inherently low assay background. |
In vivo method | Assays are performed using native chromatin, eliminating crosslinking artifacts. |
Antibody versatility | Compatible with rabbit and mouse antibodies. |
Target versatility | Generate sequencing and/or qPCR data for histones, histone modifications, transcription factors and cofactors.
|
Better Data with an Improved Protocol
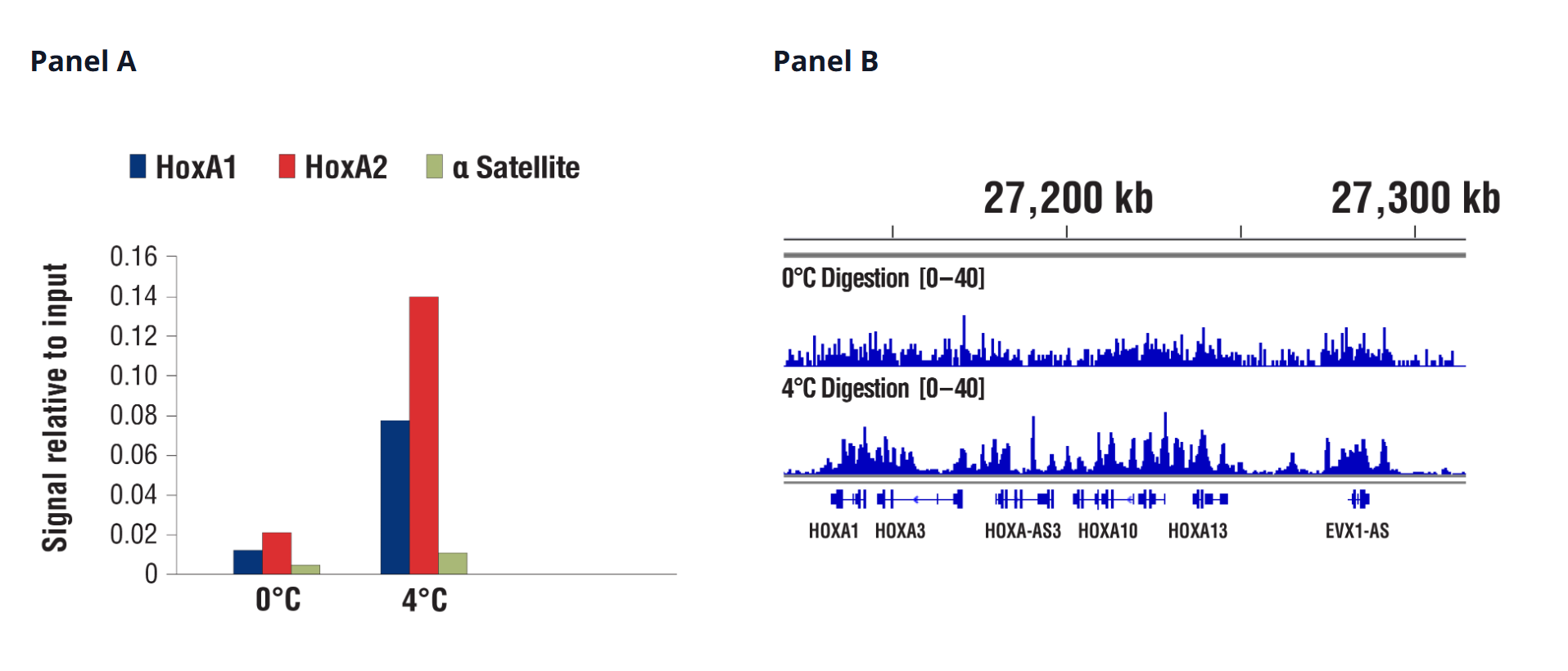
Additional development of the CST® CUT&RUN protocol led to refinements that enhance data quality, and validated both CUT&RUN Assay Kits for use with a wider variety of sample types. For example, we recommend performing digestions at 4°C instead of 0°C to facilitate the recovery of targeted chromatin fragments without significantly increasing background. This improves qPCR and NG-seq analysis, especially for transcription factors and cofactors.
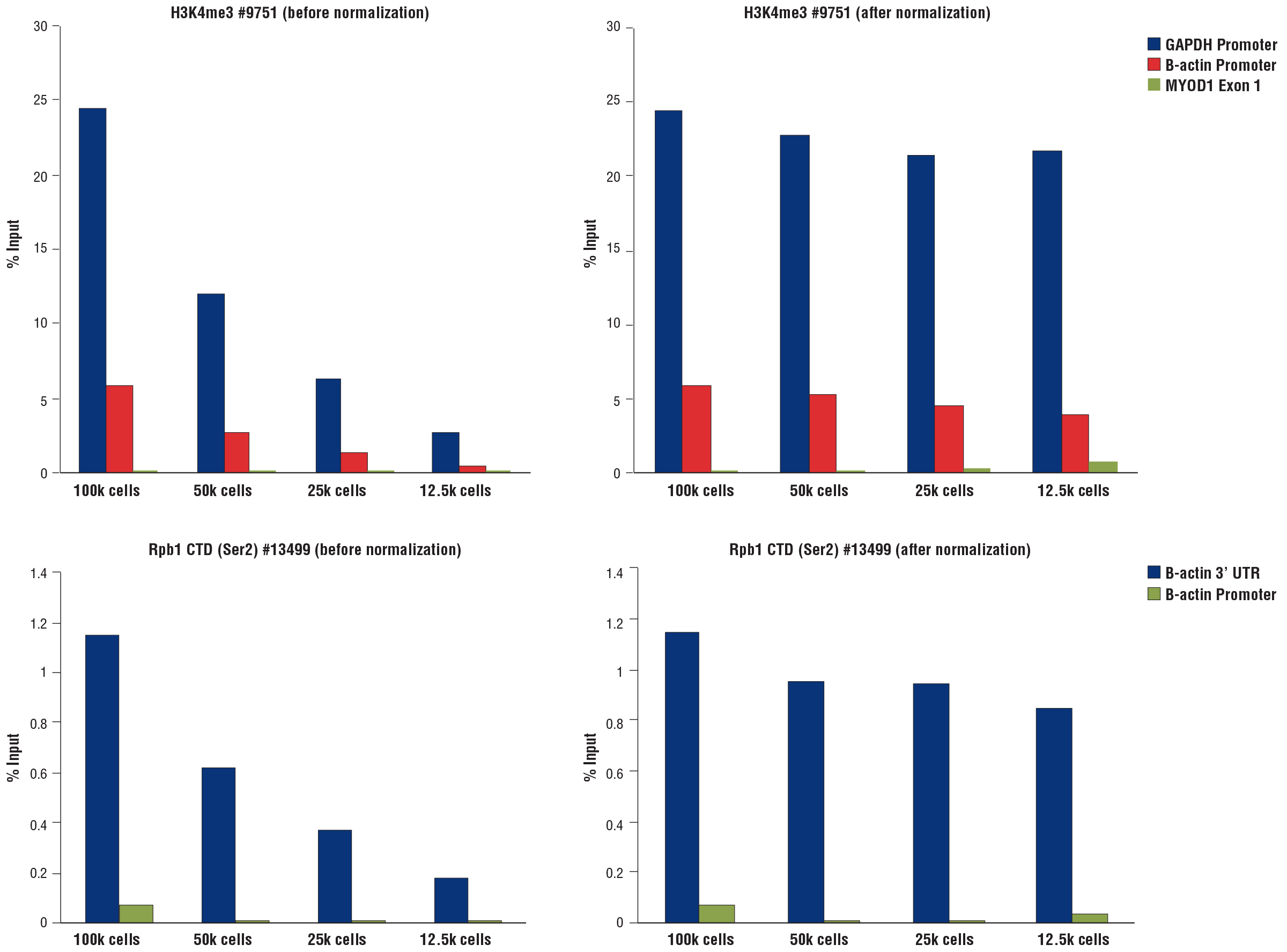
Save Time by Using 5-20x Less Sample
When available, it is recommended to use 100,000 cells for each CUT&RUN assay. However, CST realizes that this is not always possible, and has validated both our CUT&RUN kits for use with just 5,000-20,000 cells, depending on target type.
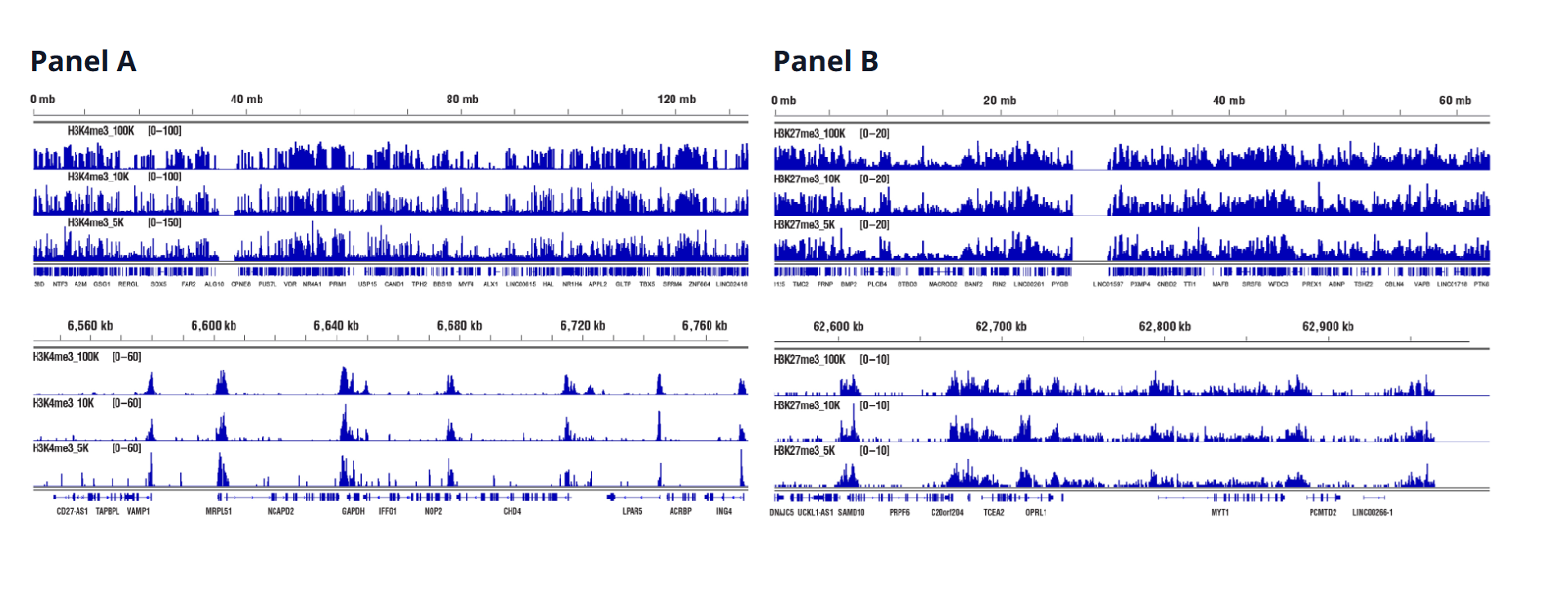
Histones (Target Versatility)
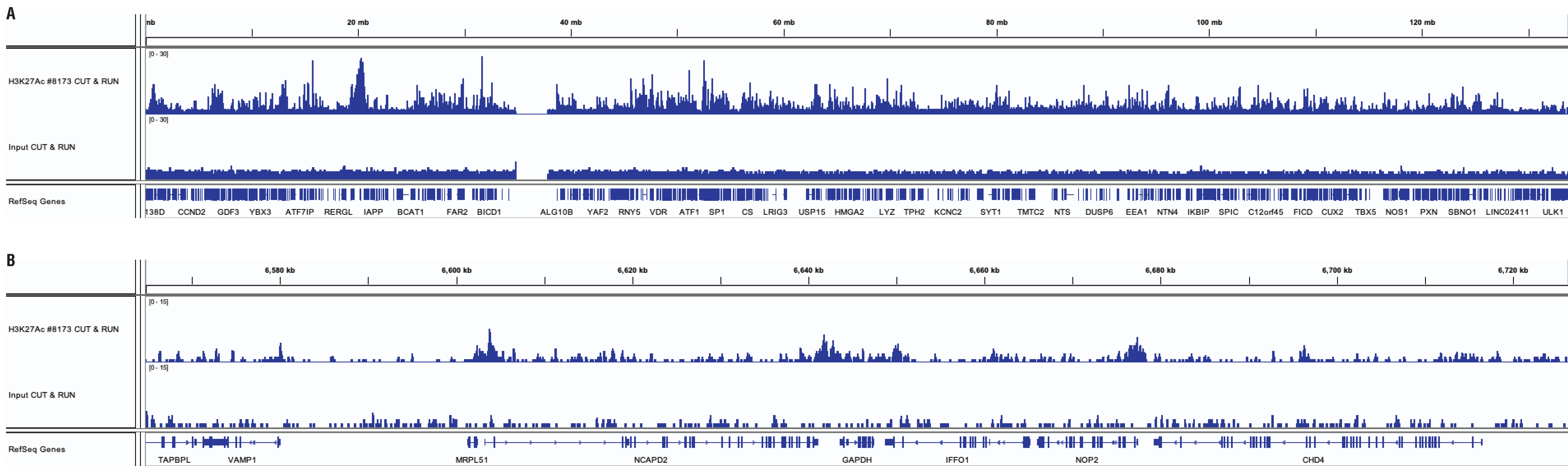
Similar binding patterns were observed when 100,000, 10,000, or 5,000 cells were used on the CUT&RUN Assay Kit #86652 when studying histones.
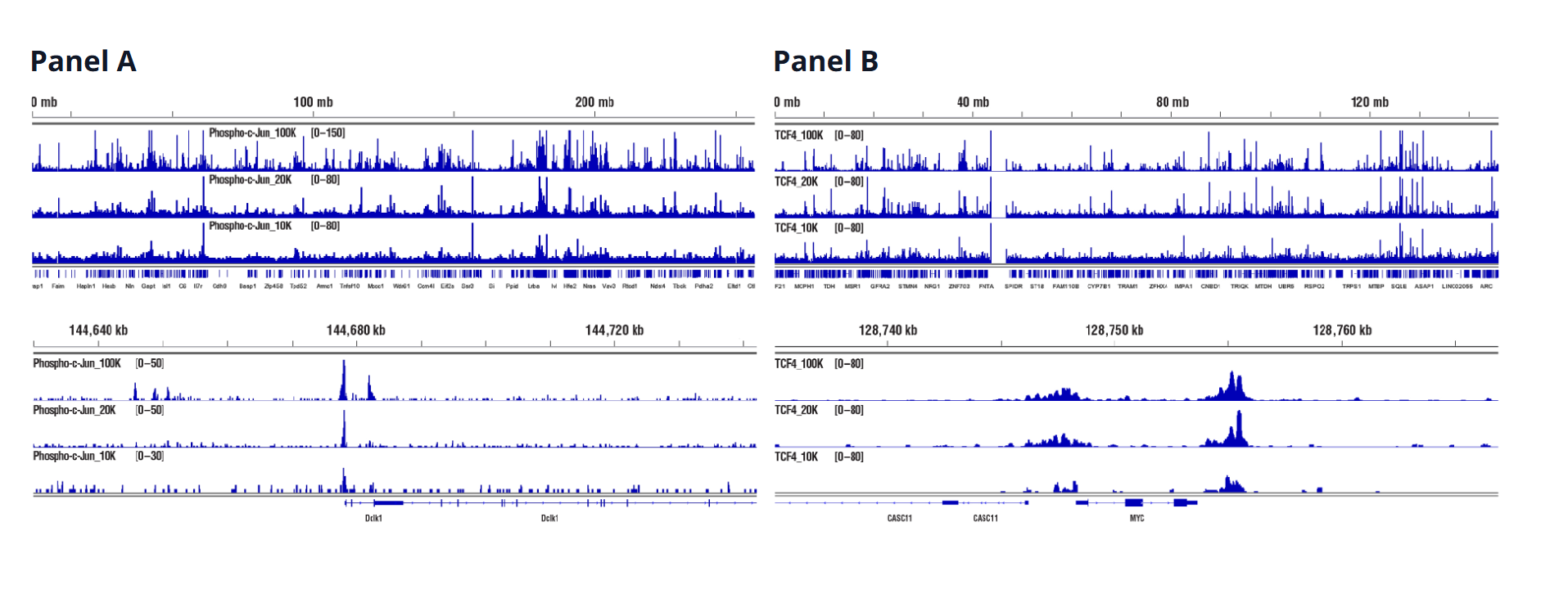
Transcription Factors (Target Versatility)
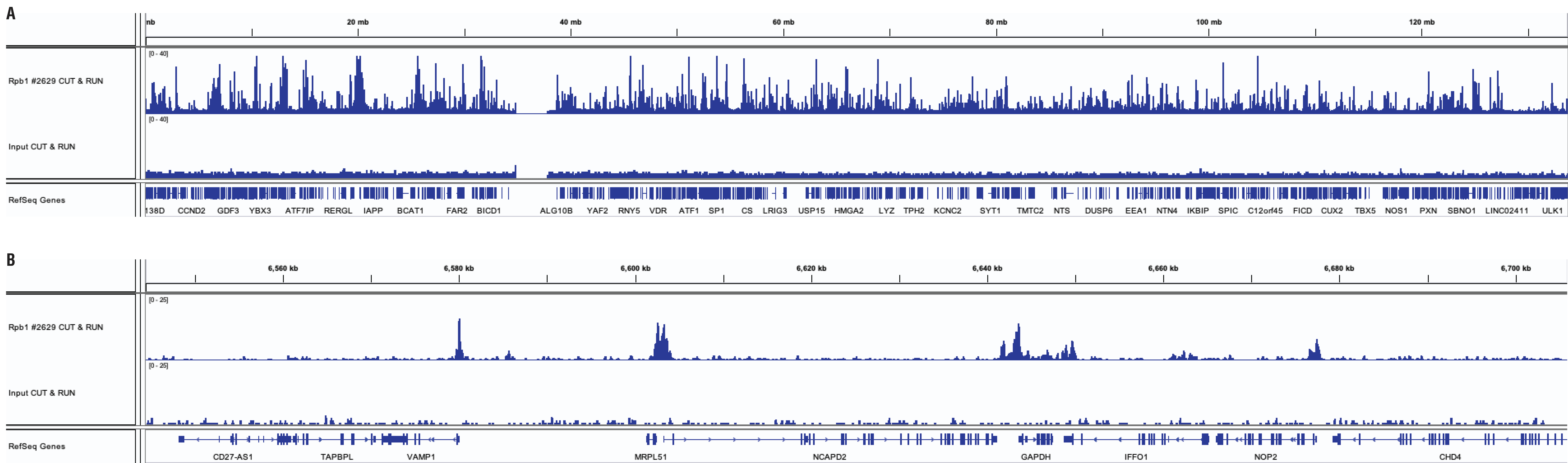
Similar binding patterns were observed when 100,000, 20,000, or 10,000 cells were used on the CUT&RUN Assay Kit #86652 when studying transcription factors.
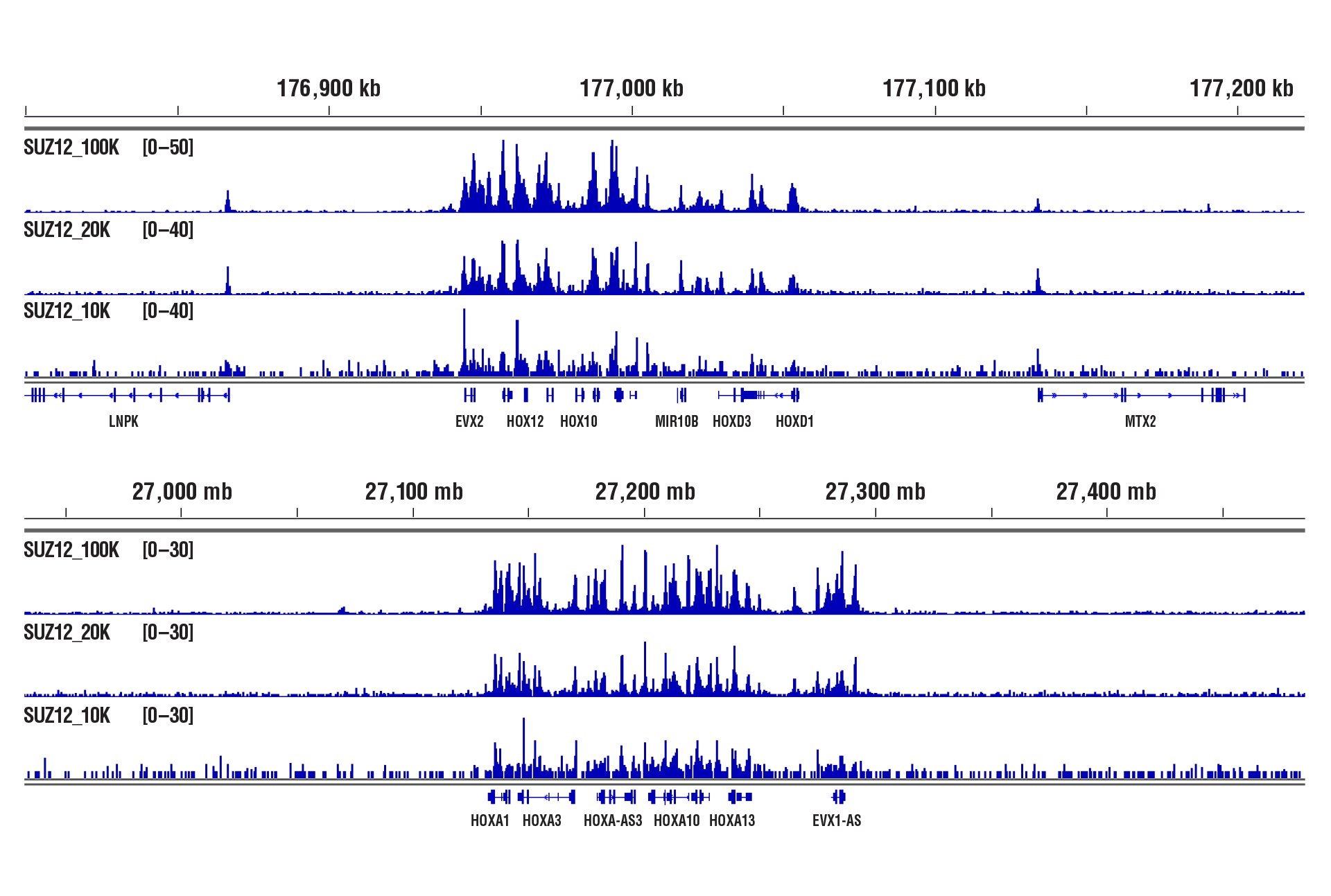
Cofactors (Target Versatility)
Similar binding patterns were observed when 100,000, 20,000, or 10,000 cells were used on the CUT&RUN Assay Kit #86652 when studying cofactors.
Study Protein-DNA Interactions in Tissue Samples with Confidence
The high sample input required to perform ChIP experiments is extremely restrictive when it comes to profiling chromatin in limited sample types like tissue. The sample savings CUT&RUN delivers for cultured cells extends to tissue samples. Depending on the tissue, ChIP can require anywhere from 25 to 50 mg of tissue, while the CST CUT&RUN kits only requires approximately 1 to 2.5 mg, depending on the tissue type.
Validated protocols give you confidence in your data, even when working with ~20x less sample to explore protein-DNA interactions using CST CUT&RUN kits.
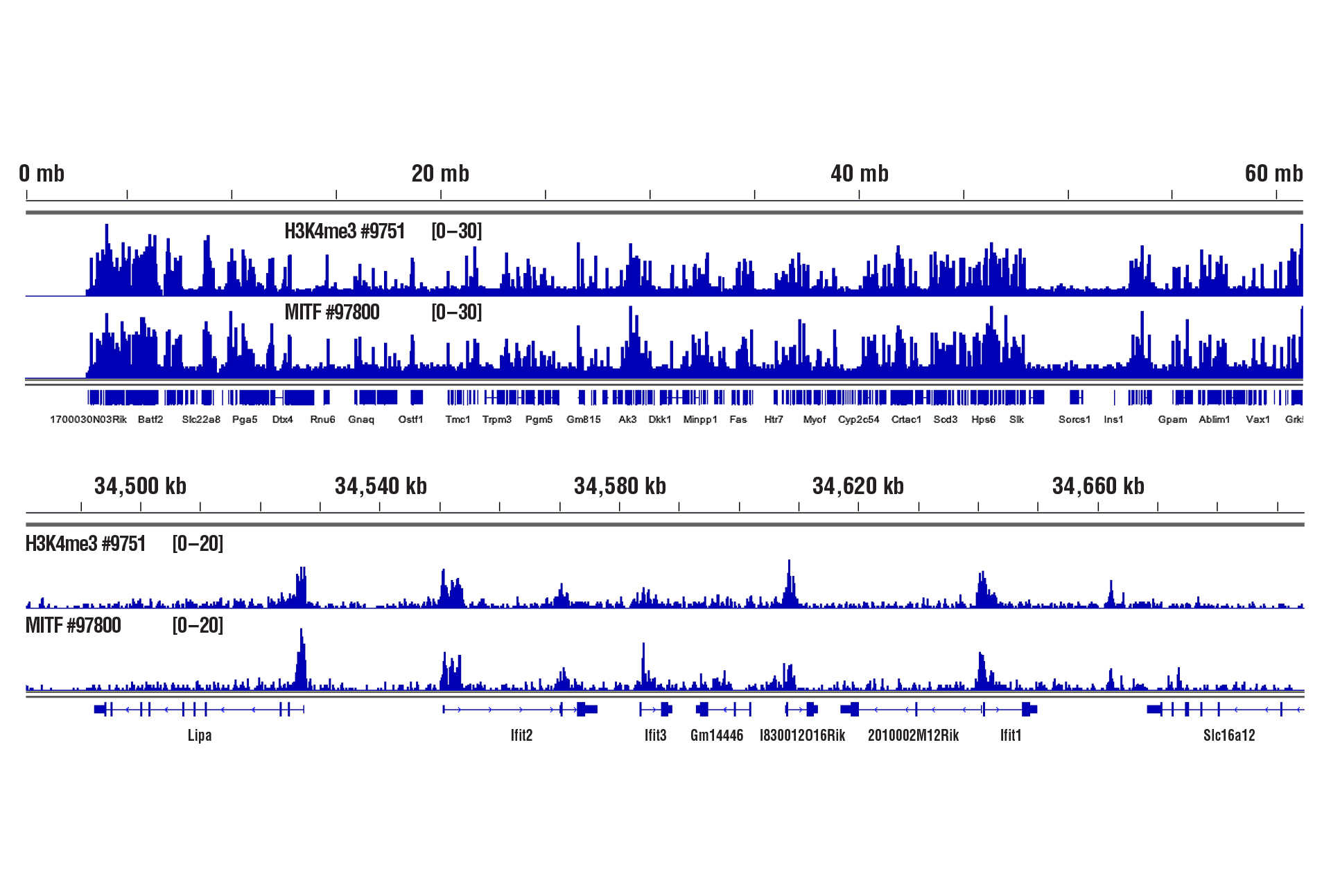
Heart Tissue – Histones and Transcription Factors
Only 1 mg of heart tissue was required to analyze protein-DNA interactions for H3K4me3 and transcription factor MITF using our CUT&RUN kits.
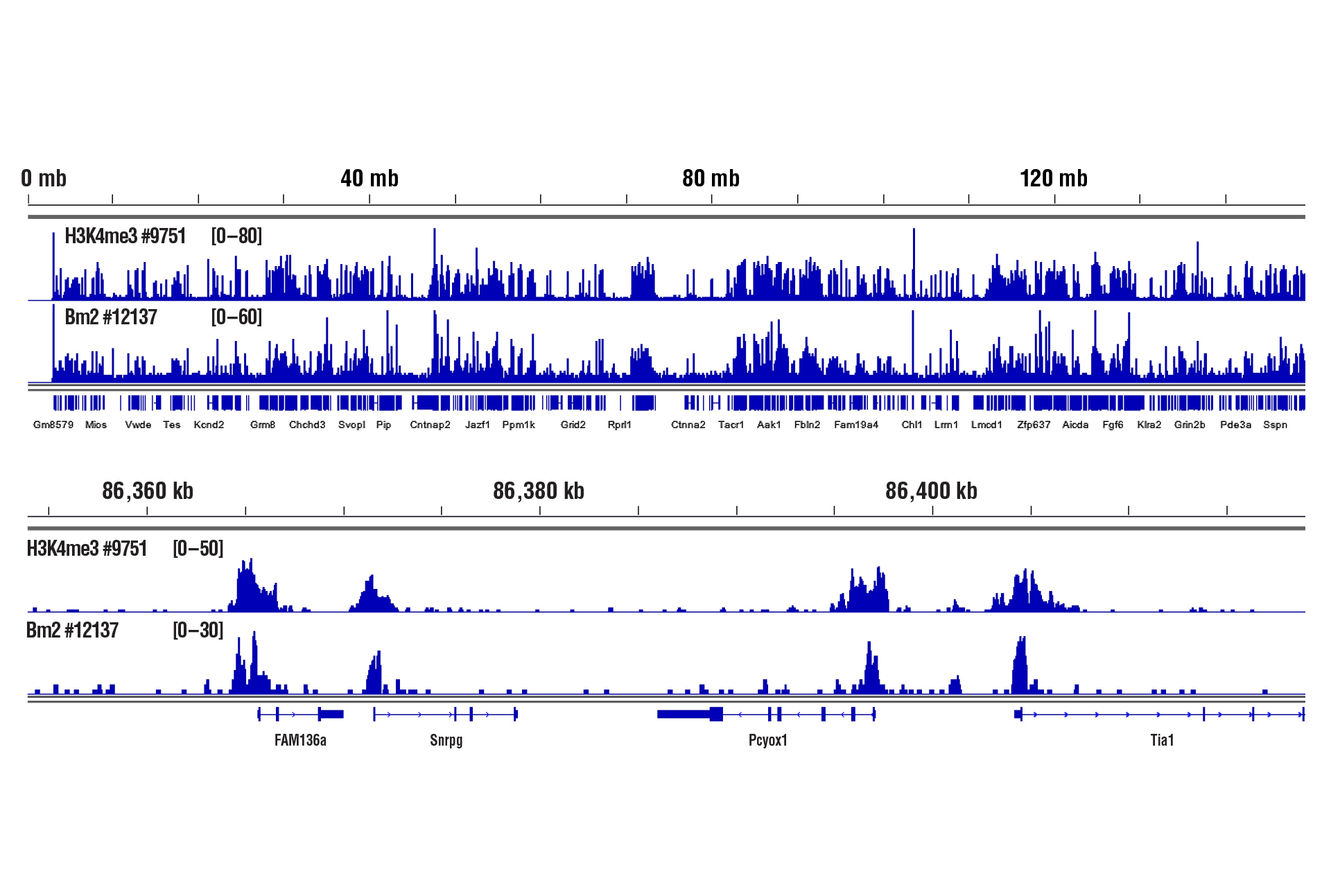
Brain Tissue – Histones and Transcription Factors
Only 1 mg of brain tissue was required to analyze protein-DNA interactions for H3K4me3 and transcription factor Brn2/POU3F2 using our CUT&RUN kits.
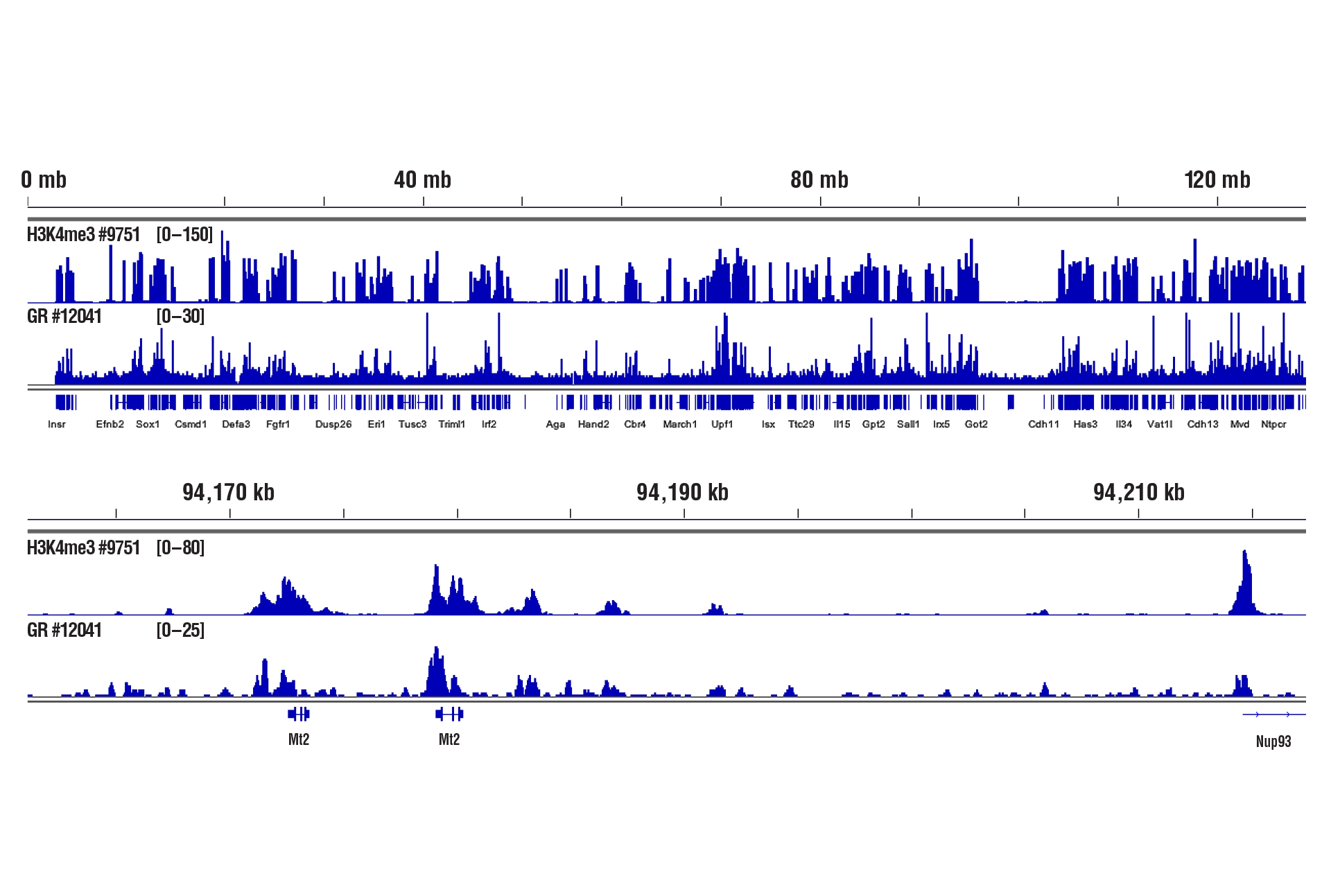
Liver Tissue – Histones and Transcription Factors
Only 1 mg of liver tissue was required to analyze protein-DNA interactions for H3K4me3 and the Glucocorticoid Receptor transcription factor using our CUT&RUN kits.
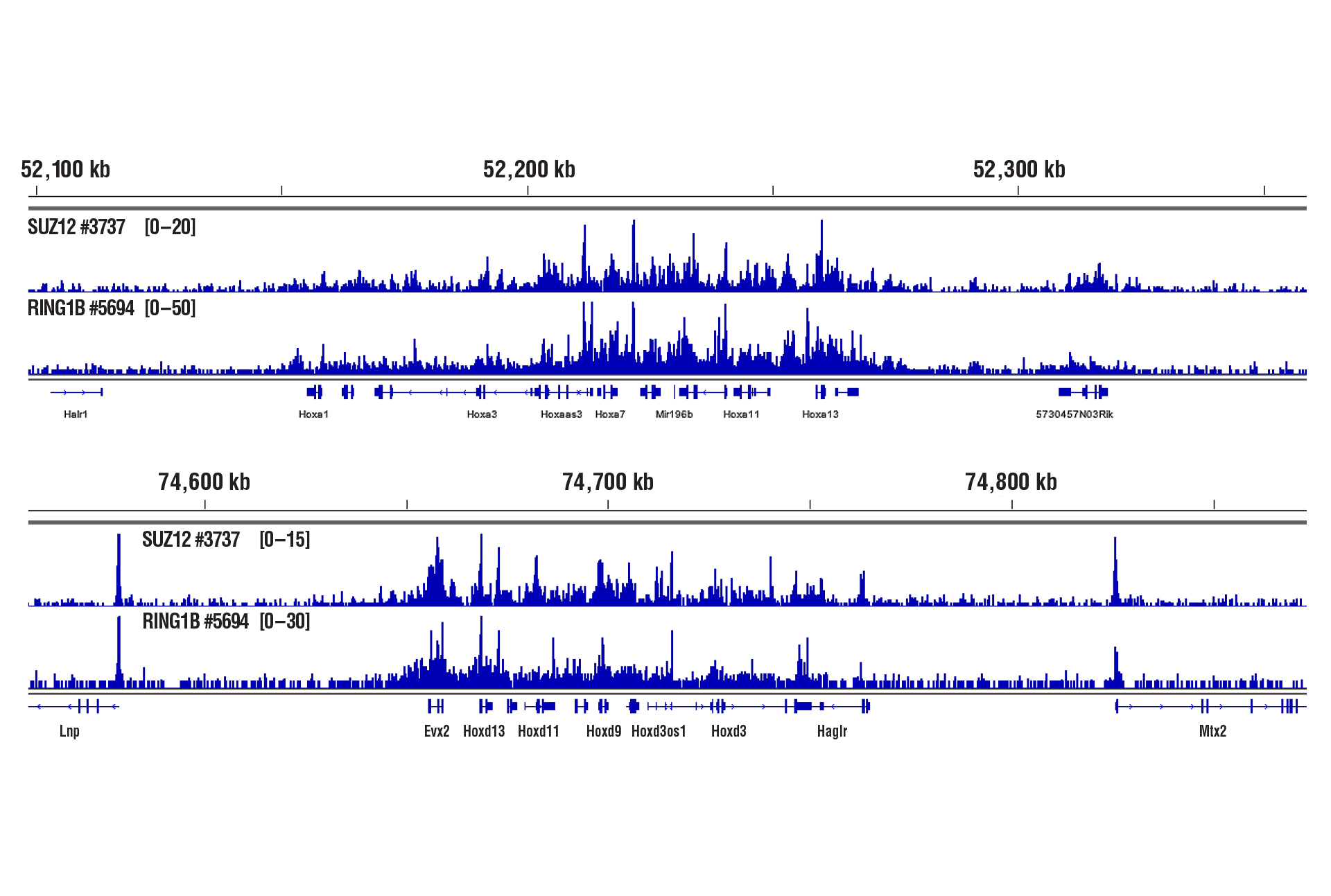
Liver Tissue – Cofactors
Only 2.5 mg of liver tissue were required to analyze protein-DNA interactions for the cofactors SUZ12 and RING1B using our CUT&RUN kits.
Profile Chromatin in Primary Cells
The CST CUT&RUN kits significantly lower the cell number requirement, making studying protein-DNA interactions in primary cells feasible. Don’t compromise your desired sample type just to meet cell number requirements.
Sequencing Cost Savings
The relatively low background means you can distinguish your signal from genomic background noise with a lower sequencing depth. Sequencing data can be obtained with just 3 to 5 million high-quality reads, reducing your sequencing costs significantly. An added bonus? The method’s low background also makes it possible to profile low-signal genomic features.
Avoid “Crosslinking Artifacts”
CUT&RUN works with native chromatin and doesn’t require crosslinking or immunoprecipitation, which can lead to “crosslinking artifacts.”
Antibody Versatility
pAG-MNase binds to both mouse and rabbit antibodies, increasing the catalog of antibodies compatible with CUT&RUN assays.
Rabbit Antibody
Mouse Antibody
Product Offerings
CST offers flexible solutions for your CUT&RUN assays, whether you’re just starting to explore gene regulation or chromatin remodeling as part of your research or you are an epigenetics expert. All you need to provide is a primary antibody against your protein of interest. CST provides everything else in two simple-to-order CUT&RUN kits depending on your spike-in control preference. Each kit contains all the buffers and reagents you need, along with a detailed protocol. Or only order the reagents you need à la carte. They are all validated in-house to ensure you’ll get the high-quality reagents and results you’ve come to expect from CST.
Catalog # | Product |
|---|---|
CUT&RUN Assay Kit (with Drosophila Spike-In Control) | |
| 86652 | CUT&RUN Assay Kit |
| 40366 | CUT&RUN pAG-MNase and Yeast Spike-in DNA |
| 14209 | DNA Purification Buffers and Spin Columns (ChIP, CUT&RUN, CUT&Tag) |
| 88989 | SimpleChIP® Universal qPCR Master Mix |
| 56795 | DNA Library Prep Kit for Illumina Systems (ChIP-seq, CUT&RUN) |
| 47538 | Multiplex Oligos for Illumina Systems (Dual Index Primers) (ChIP-seq, CUT&RUN) |
| 29580 | Multiplex Oligos for Illumina Systems (Single Index Primers) (ChIP-seq, CUT&RUN) |
| 66362 | Rabbit (DA1E) Monoclonal Antibody IgG Isotype Control (CUT&RUN) |
| 93569 | Concanavalin A Magnetic Beads and Activation Buffer |
| 16359 | Digitonin Solution |
| 27287 | 100X Spermidine |
| 7012 | Protease Inhibitor Cocktail (200X) |
| 7013 | RNAse A (10 mg/ml) |
| 10012 | Proteinase K (20mg/ml) |
| 31415 | CUT&RUN 10X Wash Buffer (CUT&RUN, CUT&Tag) |
| 15338 | Antibody Binding Buffer (CUT&RUN, CUT&Tag) |
| 48105 | CUT&RUN 4X Stop Buffer |
| 42015 | CUT&RUN DNA Extraction Buffer |
| 20533 | 10% SDS Solution |
| 7005 | Glycine Solution (10X) |
| 12606 | 16% Formaldehyde Methanol Free |
| 9872 | Phosphate Buffered Saline (PBS-1X) pH7.2 (Sterile) |
References
- Skene PJ, et al. Targeted in situ genome-wide profiling with high efficiency for low cell numbers. (2018) Nat. Protoc 13(5), 1006-1019. Pubmed 29651053
- Meers MP, et al. Improved CUT&RUN chromatin profiling and analysis tools. (2019) BioRxiv 1, 569129. bioRxiv 569129
- Skene PJ, and Henikoff S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. (2017) Elife 6, e21865. Pubmed 28079019
- Janssens DH, et al. Automated in situ chromatin profiling efficiently resolves cell types and gene regulatory programs. (2018) Epigenetics Chromatin 22(1), 74. Pubmed 30577869
- Egan, B. et al. An Alternative Approach to ChIP-Seq Normalization Enables Detection of Genome-Wide Changes in Histone H3 Lysine 27 Trimethylation upon EZH2 Inhibition. (2016) PLoS ONE 11, e0166438
- Taruttis, F. et al. External calibration with Drosophila whole-cell spike-ins delivers absolute mRNA fold changes from human RNA-Seq and qPCR data. (2017) Biotechniques 62, 53-61 Pubmed 28193148
U.S. Patent No. 11,733,248, foreign equivalents, and child patents deriving therefrom.
This product is supplied and sold under certain patents owned by Active Motif, US patents 9938524, 10689643, 11306307 and 12049622, and related patents in other countries. For purchaser’s internal research use only. May not be used for resale, services or other commercial use.
U.S. Patent No. 7,429,487, foreign equivalents, and child patents deriving therefrom.
Cell Signaling Technology and SimpleChIP are trademarks of Cell Signaling Technology, Inc.
All other trademarks are the property of their respective owners. Visit cellsignal.com/legal/trademark-information for more information.